

Standardized Centralized Alzheimer’s & Related Dementias Neuroimaging (SCAN)
The goal of SCAN is to enable standardized PET and MRI data collection from across the Alzheimer’s Disease Research Centers (ADRC) Program so that it can be combined and shared with researchers around the world via the NACC Data Platform. The National Institute on Aging (NIA) provides SCAN’s funding.
The SCAN initiative is a collaboration between:
NACC Data Release Spotlight!
SCAN PET and MRI data are now available to the scientific community.
This unprecedented dataset contains summary and analysis variables extracted from standardized amyloid PET scans, tau PET scans, and MRI scans connected to comprehensive clinical and cognitive data from NACC’s Uniform Data Set (UDS) and other data modalities available at NACC.
Submit SCAN Data
The SCAN initiative only accepts SCAN-compliant data
Images that don’t meet SCAN protocol are still accepted by NACC. Learn more about submitting mixed-protocol data to NACC.
About
Overview
The Standardized Centralized Alzheimer’s & Related Dementias Neuroimaging (SCAN) initiative is a multi-institutional project that was funded as a U24 grant (AG067418) by the National Institute on Aging (NIA) in May 2020 with the goal of standardizing the acquisition, curation, and analysis of PET and MR images acquired through the NIA Alzheimer’s Disease Research Center (ADRC) Program. The National Alzheimer’s Coordinating Center (NACC) Data Platform enriches the SCAN dataset by linking with the longitudinal Uniform Data Set and other data modalities at NACC.
ADRCs upload SCAN-compliant images to a portal hosted by the Laboratory of Neuro Imaging (LONI) at the University of Southern California where they are de-identified and defaced by the Aging and Dementia Imaging Research (ADIR) laboratory at Mayo Clinic. The PET and MRI laboratories at the University of Michigan and the Aging and Dementia Research (ADIR) Laboratory at Mayo Clinic then process for quality assurance and harmonization. Following this, the PET laboratory at UC Berkeley and the MRI laboratories at Mayo Clinic and UC Davis analyze the images to produce MRI volumes and PET Standardized Uptake Value Ratio (SUVR) data.
Recently, SCAN methods have been applied to the ADRC Consortium for Clarity in ADRD Research Through Imaging (CLARiTI).
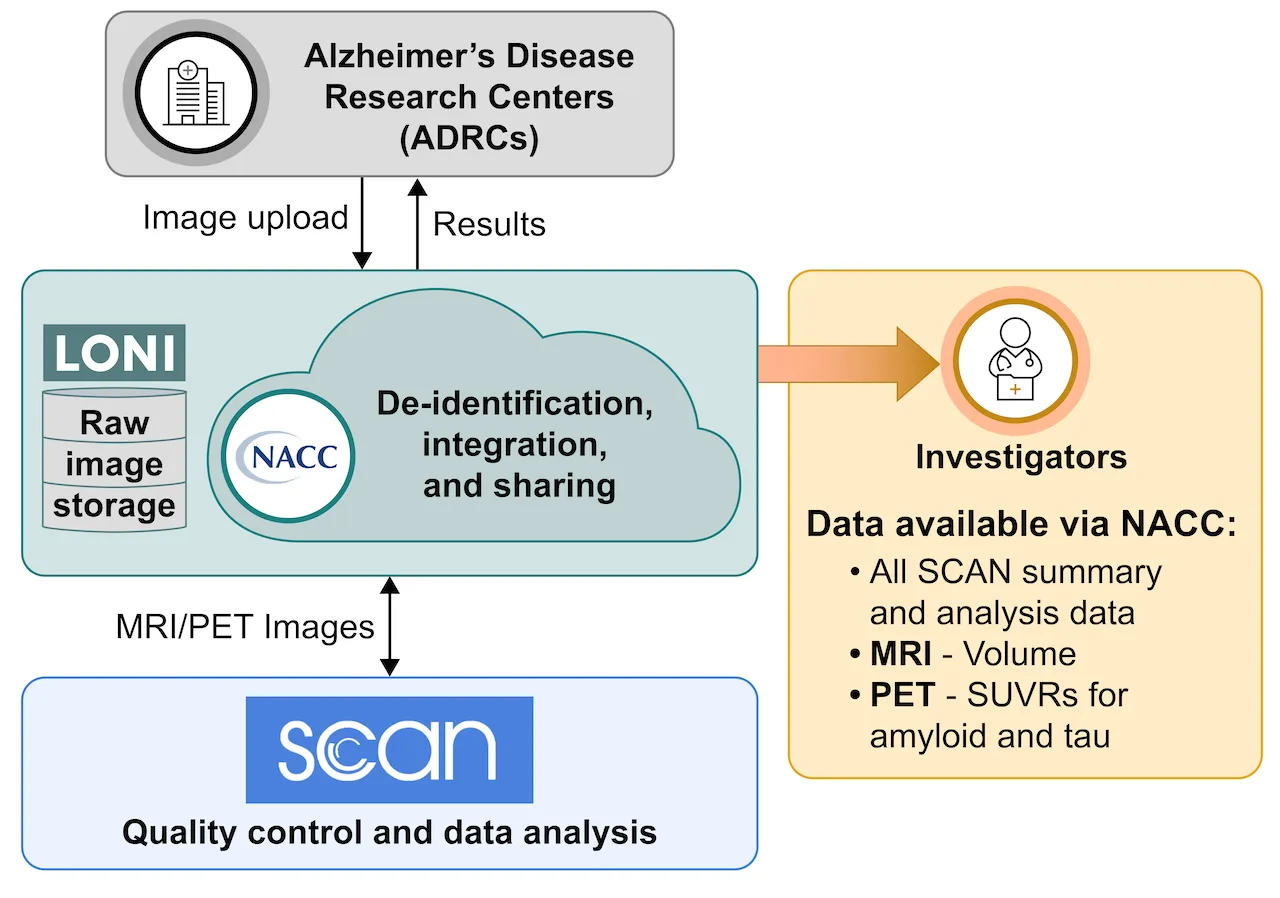
Why is SCAN needed?
For many years, researchers at the NIA-funded ADRCs have been collecting different types of PET and MR images. This “mixed protocol” or “legacy” data was collected using a variety of different acquisition methods such as T1-weighted imaging, diffusion imaging, or resting state functional MRI. Because of this, image data could not be easily combined across ADRCs, resulting in lost opportunities for scientific collaboration. The goal of SCAN is to standardize PET and MRI data collection from across the ADRC Program so that it can be combined and shared with researchers around the world via the NACC Data Platform.
Data Sharing
The NACC Data Platform makes harmonized images and numerical summaries (e.g., Standardized Uptake Value Ratio ‘SUVRs’ and volumes) available to ADRCs and approved researchers around the world through the following mechanisms:
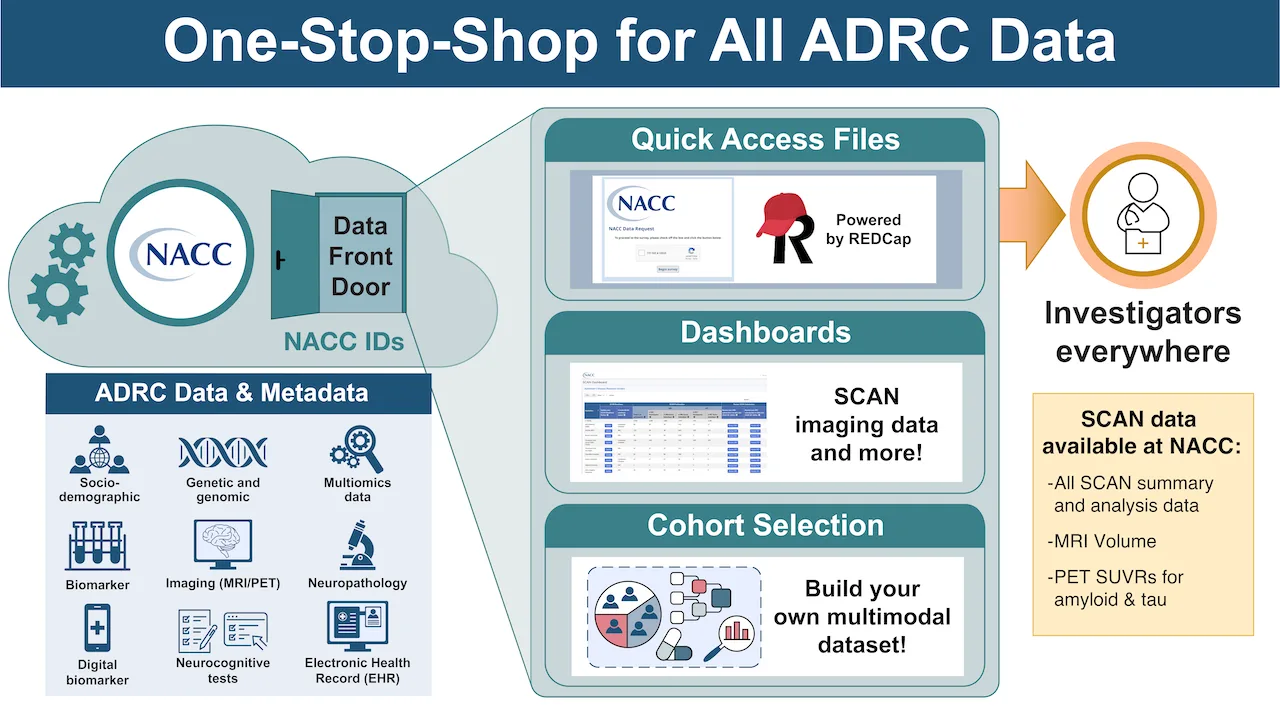
ADRC Portals: ADRCs can access and download SCAN data for their participants
SCAN QC Dashboards: ADRCs can track and audit their image submission’s quality control (QC) status and other summary information in real-time.
All SCAN Variables: Numerical analysis results – such as Standardized Uptake Value Ratio ‘SUVRs’, brain volumes, white matter hyperintensities (WMHs), cortical thickness, surface area, and cerebral infarction – are accessible to ADRCs via the ADRC Portals. Navigate to your ADRC-specific SCAN Dashboard (or ADRC Portals) to view the analysis results for all participants whose MR and PET images have passed QC and been analyzed.
NACC Data Front Door: Researchers anywhere can access SCAN data for free
SCAN Public Dashboard: The SCAN Public Dashboard provides researchers with up-to-date information on the number of participants with SCAN-compliant PET and MR images and the total number of images submitted by each ADRC to SCAN.
Quick Access File Data Request System: Researchers can request SCAN data collected from all participating ADRCs, including summary, QC, and analysis (volumes and SUVRs) data. As of August 2024, Researchers can also access defaced SCAN images via this data request system.
Cohort Selection Tool: A new self-service tool coming to NACC allowing researchers to perform cohort selection, data visualization, and provide access to raw data at NACC directly.
Raw images are stored at the Laboratory of Neuro Imaging (LONI). ADRCs and researchers interested in accessing the raw images will be able request them via LONI in the near future. NACC will provide updates on the availability of raw images as progress is made by SCAN, LONI, and NACC to ensure raw image data is de-identified before it is made available to the public.
SCAN Protocols
Prospective image files that meet SCAN acquisition protocols, regardless of the funding source, are submitted to LONI as part of this project. Existing images, and prospective images that don’t meet SCAN protocol, are instead submitted directly to NACC.
SCAN has a set of required acquisition schemes for PET and MR data. This includes required PET protocols for acquiring amyloid, tau, and fludeoxyglucose-18 (FDG) PET scans, and required MRI sequences for structural T1 and Fluid Attenuated Inversion Recovery (FLAIR) scans. Additional MRI sequences are optional. Analyzed data available to investigators will include:
Regional measures of amyloid, tau, and FDG tracer uptake
Brain volumes
White matter hyperintensities
Cerebral infarction
Fractional anisotropy, cerebral blood flow, and functional connectivity (depending on the availability of advanced MRI sequences)
Current SCAN protocols are focused on widely accepted and applied biomarkers generally, but not exclusively, related to characterizing research participants in the A/T/N framework. To continue to support collaborative research, SCAN will base future directions on the needs and goals of the ADRC program. These may include other MRI techniques or PET tracers as best practices for their acquisition and analysis become available.
SCAN Dashboard
This dashboard tracks the number of Standardized Centralized Alzheimer’s and Related Dementias Neuroimaging (SCAN)-compliant MR/PET images that have passed QC and are available for analysis. SCAN MRI and PET data that has passed QC and been analyzed is available to all researchers, worldwide, via NACC’s Data Front Door – learn more about the Data Front Door.
All data was submitted by each Alzheimer’s Disease Research Center (ADRC) to the Laboratory of Neuro Imaging (LONI) for the SCAN Initiative.
New to the SCAN Dashboards?
Showing QC-passed, analysis-ready data only
This dashboard reflects only the total number of SCAN MRI and PET data that have passed QC and are eligible for analysis. This is the data that is made available to researchers via the NACC Data Front Door – Quick Access File.
The totals reflected in the SCAN Public Dashboard may differ from the total number of images your ADRC has submitted. For a more complete picture of your total submissions by center and where they are at in the QC pipeline, please visit your ADRC-specific SCAN QC Dashboard via the “private dash” links below.
Resources for ADRCs
How to Participate
- Acquire Images
Alzheimer’s Disease Research Centers participating in SCAN should refer to the MRI and PET checklist and manuals linked below for detailed SCAN image acquisition protocols.
SCAN will troubleshoot prospectively with ADRCs to advise on making data SCAN compliant. Contact SCAN.
Current SCAN Protocols
MRI
MRI Checklist and ManualAll longitudinal scans must be performed on the same MRI scanner.
Each MRI scanner must be certified and approved by SCAN. MR scanners that are now certified for ADNI 3 can be certified for SCAN using a phantom scan. MR scanners that have not been certified for ADNI 3 (or SCAN) must be certified using a human volunteer scan. This must be permissible using a local IRB – i.e. SCAN does not plan to create a central IRB nor will SCAN manage local IRB administrative proceedures.
The Standard MRI protocol (using SCAN-specific parameters) is required. Your remaining scan time can be used for your site specific sequences.
There are 2 options for SCAN MR – each site can choose which they prefer.
Option 1: Site only required to do 2 primary SCAN sequences: T1 & FLAIRThe first 10 minutes of exam belongs to NIA/SCAN — this is when the SCAN T1 & FLAIR would be done
Remainder of exam is unique to site
SCAN would only collect and analyze T1 and FLAIR
PET
PET Checklist and ManualAll longitudinal scans must be performed on the same PET scanner.
Each PET scanner must be certified and approved by SCAN. ADNI approved PET scanners will not need to be re-certified.
The following tracers and imaging protocols are all compatible with SCAN:
Suggested Target Dose ± 10% Minimum injectable dose Acquisition start-stop time post-injection (min) Tracer mCi MBq mCi MBq Amyloid Tracers PIB 15 555 8 300 40–60 or 50–70, 40–70 Florbetapir 10 370 6 225 50–70 Florbetaben 8 295 5 185 90–110 NAV4694 8.1 300 6.5 240 40–70, 50–70 Flutemetamol 5 185 4 150 90–110 TAU Tracers Flortaucipir 10 370 6 225 80–100 MK6240 5 185 4 150 70–90 or 90–110, 70–110 PI2620 5 185 4 150 30–60, 45–75, 60–90, 45–90 GTP1 7 260 5 185 60–90 Glucose metabolism tracer FDG 5 185 4 150 30–45, 30–60 NOTE: The SCAN MR protocol is the same as the Alzheimer’s Disease Neuroimaging Initiative (ADNI) protocol. The current protocol is ADNI 4. See this set of ADNI MR protocols created for various vendor/model/operating system options.
- Upload Images
Submit SCAN-Compliant data to LONI
Please carefully review the SCAN PET manual and the SCAN MRI manual to ensure that your data is compliant with the SCAN protocol and for uploading instructions.
Only SCAN-compliant data should be submitted to the Laboratory of Neuro Imaging (LONI). Non-approved data (with or without approved data) will be quarantined. Images collected before 2021 do not meet SCAN-compliance criteria.
Submit non–SCAN-compliant data directly to NACC
SCAN does not accept data that is not prospectively collected and SCAN-compliant.
Any data that that is 'mixed protocol' or non-SCAN-compliant should be submitted to NACC.
- Track and audit image submissions
Sites are responsible for auditing their submissions and making sure that the information is correct using the SCAN QC Dashboards.
Each ADRC can access their ADRC-specific SCAN QC Dashboards via the SCAN Public Dashboard.
These enable sites to track the QC status of their MRI and PET submissions, allowing them to rapidly address errors. They also serve as an auditable record of all SCAN MRI data submitted to SCAN by a given ADRC and are downloadable as a csv file.
NOTE: The ADRC-specific SCAN QC Dashboards are only available to ADRC staff with data access privileges. Please refer to the SCAN Dashboard Quick Access Guide for details.
- Access your SCAN data and analysis results
All SCAN variables will soon be available via ADRC Portals. We anticipate this data being available by Fall 2024.
In the meantime, ADRCs may access analysis results by submitting a data request via NACC’s Quick Access File Data Request System.
NOTE: Analysis results are calculated after defacing and QC has been performed on MR and PET images. Sites should expect a delay from when images are submitted to when analysis results are available.
Data Resources
See BelowSCAN team Tips for successful uploads
Qualify all scanner(s) to the SCAN protocol for MRI and PET as per the SCAN MRI manual and the SCAN PET manual.
- For MRI: Contact the Mayo ADIR Lab at scanmri@mayo.edu
- For PET: Contact Suzanne Baker at slbaker@lbl.gov
Do not self-deface series or scans.
- This is done centrally by SCAN for all relevant series prior to image release.
- Defacing multiple times introduces undesirable data heterogeneity.
Do not self-anonymize DICOMs.
- This is done by LONI upon image upload.
- May be impossible to protocol check or even display an image.
Do not upload the same exam(s) or scan(s) multiple times.
- Uploading exams multiple times results in unnecessary investigation and clean up time/effort which diminishes throughput.
Ensure correct participant ID is assigned to the upload exam.
- Failure to do so results in unnecessary investigation and clean up time/effort which diminishes throughput.
For PET, do not upload non-attenuation corrected data (NAC), also do not upload summed frames (anything longer than 5 min per frame).
Resources for Researchers
Researchers around the world can access Standardized Centralized Alzheimer’s Disease and Related Dementias Neuroimaging (SCAN) initiative harmonized images and numerical summaries (e.g. Standardized uptake value ratio ‘SUVRs’ and volumes) via the National Alzheimer’s Coordinating Center (NACC) Data Platform through NACC’s Data Front Door.
NACC's Data Front Door
Free access is available via the following mechanisms:
SCAN Public Dashboard
Up-to-date information about the number of participants with SCAN-compliant MR/PET images and the total number of images submitted by each Alzheimer’s Disease Research Center (ADRC) to SCAN.
Quick Access File Data Request System
Researchers can request SCAN numerical data collected from all participating ADRCs, including summary, QC, and analysis (volumes, cortical thickness, surface area, and SUVRs) data. As of August 2024, Researchers can also access defaced SCAN images via this data request system.
Raw Image Availability
Images are currently stored at the Laboratory of Neuro Imaging (LONI). Researchers can access these images via NACC’s Quick Access File Data Request System. NOTE: Due to the processing time needed to perform defacing, QC, and analysis on submitted data, not all image data that has been submitted and/or passed through QC is available to researchers at this time.
Cohort Selection Tool:
A new self-service tool coming to NACC allowing researchers to perform cohort selection, data visualization, and provide access to raw data at NACC directly.
Data Acknowledgements
IF USING MRI AND/OR PET DATA FROM THE STANDARDIZED CENTRALIZED ALZHEIMER’S DISEASE NEUROIMAGING (SCAN) INITIATIVE, A STATEMENT ACKNOWLEDGING THE SCAN GRANT AND CENTERS IS REQUIRED IN PRESENTATIONS AND THE ACKNOWLEDGMENTS SECTION OF MANUSCRIPTS AS DETAILED IN THE NACC publication guidelines.
Data Resources
- SCAN MRI Researcher’s Data Dictionary
- SCAN PET Researcher’s Data Dictionary
- SCAN MRI QC AND ANALYSIS Methods Documentation
- SCAN MRI Analysis Methods Documentation
- Tau PET (MRI-free) Methods Documentation
- Amyloid PET (MRI-free) Methods Documentation
- FDG PET (MRI-free) Methods Documentation
Data Acknowledgements
IF USING MRI AND/OR PET DATA FROM THE STANDARDIZED CENTRALIZED ALZHEIMER’S DISEASE NEUROIMAGING (SCAN) INITIATIVE, A STATEMENT ACKNOWLEDGING THE SCAN GRANT AND CENTERS IS REQUIRED IN PRESENTATIONS AND THE ACKNOWLEDGMENTS SECTION OF MANUSCRIPTS AS DETAILED IN THE NACC publication guidelines.
Meet the Team
Standardized Centralized Alzheimer’s and Related Dementias Neuroimaging (SCAN) Initiative Team
MRI Team
Clifford Jack, MD - Mayo Clinic (Principal Investigator)
Charles DeCarli, MD
Chadwick Ward
Pauline Maillard
Christopher Schwarz
Denise Reyes
Bret Borowski
John Moore-Weiss
Leonard Matoush
Robert Reid
Anne Effron
Gregory Preboske
Jeffrey Gunter
Matthew Senjem
Colin Hortman
Kejal Kantarci
Oliver Martinez
PET Team
William Jagust, MD - UC Berkeley (Principal Investigator)
Bob Koeppe
Suzanne Baker
Tessa Harrison
Trevor Chadwick
LONI Team
Arthur Toga
Karen Crawford
NACC Team
Heather O'Connell
Jessica Culhane
Zach Stark
Brendan Smith
SCAN Updates
SCAN MR and PET Image Availability
As of August 2024, researchers can now request defaced SCAN images via NACC's Quick Access File Data Request System, in addition to the SCAN numerical data collected from all participating ADRCs which includes summary, QC, and analysis (volumes, cortical thickness, surface area, and SUVRs) data.
SCAN MRI and PET Data is submitted by several Alzheimer’s Disease Researcher Centers (ADRCs) following clearly defined SCAN MRI and PET acquisition protocols. All SCAN image data has been collected and submitted after January 2021.
Documentation, such as the SCAN MRI and PET Data Dictionaries can be found in the Data Resources section above.
NOTE: Due to the processing time needed to perform defacing, not all numerical SCAN data will have associated MR or PET image data available to researchers at this time.
PET Summary and Analysis Data Availability
In January 2024, SCAN PET summary and analysis (SUVRs) data was returned to ADRCs that have contributed to SCAN. Sites were sent a summary report of the PET data each site has uploaded to SCAN, in addition to the PET numerical data analysis report which contains the derived outputs of the PET image data after it has passed through the SCAN PET QC pipeline, which includes QC of the incoming images as well as QC inspection of the products of the analytic pipelines.
The PET data file contained the following:
PET summary and QC data
PET SUVRs
See the SCAN PET Researcher’s Data Dictionary for documentation, methods, and related materials.
NOTE: Due to the processing time needed to perform defacing, QC, and analysis on submitted data, not all data that has been submitted and/or passed through QC has analysis data associated with it.
MRI Summary and Analysis Data Availability
In March 2024, SCAN MRI summary and analysis (analysis) data was returned to ADRCs that have contributed MRI data to SCAN. Sites were sent a summary report of the MRI data each site has uploaded to SCAN, in addition to the MRI volumetric analysis report which contains the derived outputs of the MRI image data after it has passed through the SCAN MRI QC pipeline.
The MRI data file contained the following:
MRI summary and QC data
MRI brain volumes, cortical thickness, and surface area data
About the SCAN analysis process
The SCAN MR analysis team performs standard processing of all MR images that have passed QC using FreeSurfer to provide standard readouts. These include the parcellations of regional brain volumes, cortical thickness and surface area for 31 bilateral gray matter regions from the Desikan-Killiany atlas.
NOTE: Due to the processing time needed to perform defacing, QC, and analysis on submitted data, not all data that has been submitted and/or passed through QC has analysis data associated with it.
See the SCAN MRI Researcher’s Data Dictionary for documentation, methods, and related materials.
The LONI uploader includes a new verification step
Beginning July 29, 2023 the LONI imaging data submission interface will require that sites enter both the PTID and the NACCID for a given participant to submit their MRI/PET data to SCAN. This change was introduced to ensure NACCID and data fidelity for the SCAN initiative. As shared in our ADRC-wide communication on March 17, 2023, ADRCs are currently only able to submit SCAN data for participants with existing NACCIDs. More details about this temporary limitation and next steps can be found in the What is the current process for submitting MRI/PET images? section of this page.How to correctly submit your data via the new LONI uploader
Sites must enter both identifiers into the Subject ID text field using the convention PTID+NACCID, where PTID and NACCID are separated by the
+character.If the given PTID and NACCID pair match the record present in the NACC database, the upload process will be allowed to continue.
If not, then the user will receive an error message indicating that the PTID and NACCID combination are not valid, and the upload will not be allowed to continue.
Users who do not follow the PTID+NACCID convention will see a message describing the expected formatting of the Subject ID.
How do I find the NACCIDs for my participants?
To access your site’s PTID and NACCID pairs, please work with your site’s data manager to utilize the PTID to NACCID Map tool available via the NACC portal.
Publications using the SCAN Initiative
With each data file request NACC records a "proposal", including a brief description of the project being undertaken. These proposals can lead to one or more publications. All resulting publications can be found in the list below.
SCAN Bibliography
- Cody KA, Sokołowski A, Johns E, Guerra LM, Winer JR, Young CB, Younes K, Dumitrescu L, Archer DB, Durant A, Sathe A, Koran MEI, Mez J, Saykin AJ, Toga A, Cuccaro M, Tosun D, Insel PS, Johnson SC, Harrison TM, Hohman TJ, Mormino EC. Characterizing amyloid and tau positron emission tomography-based stages across the clinical continuum. Alzheimers Dement. 2026 Jan;22(1):e71017. doi: 10.1002/alz.71017. PMID: 41505234; PMCID: PMC12782155. https://doi.org/10.1002/alz.71017
- Venkatesh R, Cardone KM, Bradford Y, Moore AK, Kumar R, Moore JH, Shen L, Kim D, Ritchie MD. Integrative multi-omics approaches identify molecular pathways and improve Alzheimer's Disease risk prediction. Alzheimers Dement. 2025 Nov;21(11):e70886. doi: 10.1002/alz.70886. PMID: 41231230; PMCID: PMC12614089. https://doi.org/10.1002/alz.70886
- Cody KA, Sokolowski A, Guerra LM, Insel PS, Jagust WJ, Harrison TM, Mormino EC. Comparison of amyloid PET acquired through standardized and unstandardized protocols. Alzheimers Dement. 2025 Sep;21(9):e70697. doi: 10.1002/alz.70697. PMID: 40983958; PMCID: PMC12453954. https://doi.org/10.1002/alz.70697
- Kuzma A, Valladares O, Greenfest-Allen E, Nicaretta H, Kirsch M, Ren Y, Katanic Z, White H, Wilk A, Bass L, Brettschneider J, Carter L, Cifello J, Chuang WH, Clark K, Gangadharan P, Haut J, Ho PC, Horng W, Iqbal T, Jin Y, Keskinen P, Rose AL, Moon MK, Manuel J, Qu L, Robbins F, Saravanan N, Sha J, Tate S, Zhao Y; Alzheimer's Disease Sequencing Project; Cantwell L, Gardner J, Chou SY, Tzeng JY, Bush W, Naj A, Kuksa P, Lee WP, Leung YY, Schellenberg G, Wang LS. NIAGADS: A data repository for Alzheimer's disease and related dementia genomics. Alzheimers Dement. 2025 Jun;21(6):e70255. doi: 10.1002/alz.70255. PMID: 40545618; PMCID: PMC12183107. https://doi.org/10.1002/alz.70255
- Kang K, Zhang P, Dumitrescu L, Mukherjee S, Lee ML, Choi SE, Trittschuh EH, Mez J, Saykin AJ, Gifford KA, Buckley RF, Gao X, Di J, Crane PK, Hohman TJ, Liu D. The dynamics of cognitive decline toward Alzheimer's disease progression: Results from ADSP-PHC's harmonized cognitive composites. Alzheimers Dement. 2025 Jun;21(6):e70335. doi: 10.1002/alz.70335. PMID: 40538025; PMCID: PMC12179338. https://doi.org/10.1002/alz.70335
- Younes K, Johns E, Young CB, Kennedy G, Mukherjee S, Vossler HA, Winer J, Cody K, Henderson VW, Poston KL, Betthauser TJ, Bevis B, Brooks WM, Burns JM, Coombes SA, DeCarli C, DiFilippo FP, Duara R, Fan AP, Gibbons LE, Golde T, Johnson SC, Lepping RJ, Leverenz J, McDougall S, Rogalski E, Sanders E, Pasaye J, Sridhar J, Saykin AJ, Sridharan A, Swerdlow R, Trittschuh EH, Vaillancourt D, Vidoni E, Wang WE, Mez J, Hohman TJ, Tosun D, Biber S, Kukull WA, Crane PK, Mormino EC. Amyloid PET predicts longitudinal functional and cognitive trajectories in a heterogeneous cohort.Alzheimers Dement. 2025 Mar;21(3):e70075. doi: 10.1002/alz.70075. PMID: 40145384; PMCID: PMC11947745. https://doi.org/10.1002/alz.70075
- Landau SM, Harrison TM, Baker SL, Boswell MS, Lee J, Taggett J, Ward TJ, Chadwick T, Murphy A, DeCarli C, Schwarz CG, Vemuri P, Jack CR Jr, Koeppe RA, Jagust WJ; U.S. POINTER Study Group and for the Alzheimer's Disease Neuroimaging Initiative. Positron emission tomography harmonization in the Alzheimer's Disease Neuroimaging Initiative: A scalable and rigorous approach to multisite amyloid and tau quantification. Alzheimers Dement. 2025 Jan;21(1):e14378. doi: 10.1002/alz.14378. Epub 2024 Nov 19. PMID: 39559932; PMCID: PMC11772732. https://doi.org/10.1002/alz.14378
- Johns E, Vossler HA, Young CB, Carlson ML, Winer JR, Younes K, Park J, Rathmann-Bloch J, Smith V, Harrison TM, Landau S, Henderson V, Wagner A, Sha SJ, Zeineh M, Zaharchuk G, Poston KL, Davidzon GA, Mormino EC. Florbetaben amyloid PET acquisition time: Influence on Centiloids and interpretation. Alzheimers Dement. 2024 Aug;20(8):5299-5310. doi: 10.1002/alz.13893. Epub 2024 Jul 4. PMID: 38962867; PMCID: PMC11350032. https://doi.org/10.1002/alz.13893
- Pezzoli S, Giorgio J, Martersteck A, Dobyns L, Harrison TM, Jagust WJ. Successful cognitive aging is associated with thicker anterior cingulate cortex and lower tau deposition compared to typical aging. Alzheimers Dement. 2024 Jan;20(1):341-355. doi: 10.1002/alz.13438. Epub 2023 Aug 24. PMID: 37614157; PMCID: PMC10916939. https://doi.org/10.1002/alz.13438
- Younes K, Mormino EC. The pathotome and precision health. Brain. 2023 Jun 1;146(6):2208-2210. doi: 10.1093/brain/awad154. PMID: 37150902; PMCID: PMC10232233. https://doi.org/10.1093/brain/awad154
- Weiner MW, Veitch DP, Miller MJ, Aisen PS, Albala B, Beckett LA, Green RC, Harvey D, Jack CR Jr, Jagust W, Landau SM, Morris JC, Nosheny R, Okonkwo OC, Perrin RJ, Petersen RC, Rivera-Mindt M, Saykin AJ, Shaw LM, Toga AW, Tosun D, Trojanowski JQ; Alzheimer's Disease Neuroimaging Initiative. Increasing participant diversity in AD research: Plans for digital screening, blood testing, and a community-engaged approach in the Alzheimer's Disease Neuroimaging Initiative 4. Alzheimers Dement. 2023 Jan;19(1):307-317. doi: 10.1002/alz.12797. Epub 2022 Oct 9. PMID: 36209495; PMCID: PMC10042173. https://doi.org/10.1002/alz.12797
- Tosun D, Demir Z, Veitch DP, Weintraub D, Aisen P, Jack CR Jr, Jagust WJ, Petersen RC, Saykin AJ, Shaw LM, Trojanowski JQ, Weiner MW; Alzheimer's Disease Neuroimaging Initiative. Contribution of Alzheimer's biomarkers and risk factors to cognitive impairment and decline across the Alzheimer's disease continuum.Alzheimers Dement. 2022 Jul;18(7):1370-1382. doi: 10.1002/alz.12480. Epub 2021 Oct 14. PMID: 34647694; PMCID: PMC9014819. https://doi.org/10.1002/alz.12480
- Veitch DP, Weiner MW, Aisen PS, Beckett LA, DeCarli C, Green RC, Harvey D, Jack CR Jr, Jagust W, Landau SM, Morris JC, Okonkwo O, Perrin RJ, Petersen RC, Rivera-Mindt M, Saykin AJ, Shaw LM, Toga AW, Tosun D, Trojanowski JQ; Alzheimer's Disease Neuroimaging Initiative. Using the Alzheimer's Disease Neuroimaging Initiative to improve early detection, diagnosis, and treatment of Alzheimer's disease. Alzheimers Dement. 2022 Apr;18(4):824-857. doi: 10.1002/alz.12422. Epub 2021 Sep 28. PMID: 34581485; PMCID: PMC9158456. https://doi.org/10.1002/alz.12422
- Hampel H, Cummings J, Blennow K, Gao P, Jack CR Jr, Vergallo A. Developing the ATX(N) classification for use across the Alzheimer disease continuum. Nat Rev Neurol. 2021 Sep;17(9):580-589. doi: 10.1038/s41582-021-00520-w. Epub 2021 Jul 8. PMID: 34239130. https://doi.org/10.1038/s41582-021-00520-w
- Knopman DS, Jagust WJ. Alzheimer Disease Spectrum: Syndrome and Etiology From Clinical and PET Imaging Perspectives. Neurology. 2021 Feb 16;96(7):299-300. doi: 10.1212/WNL.0000000000011415. Epub 2020 Dec 22. PMID: 33361254; PMCID: PMC8570920. https://doi.org/10.1212/WNL.0000000000011415
- Vasanthakumar A, Davis JW, Idler K, Waring JF, Asque E, Riley-Gillis B, Grosskurth S, Srivastava G, Kim S, Nho K, Nudelman KNH, Faber K, Sun Y, Foroud TM, Estrada K, Apostolova LG, Li QS, Saykin AJ; Alzheimer's Disease Neuroimaging Initiative (ADNI). Harnessing peripheral DNA methylation differences in the Alzheimer's Disease Neuroimaging Initiative (ADNI) to reveal novel biomarkers of disease. Clin Epigenetics. 2020 Jun 15;12(1):84. doi: 10.1186/s13148-020-00864-y. PMID: 32539856; PMCID: PMC7294637. https://doi.org/10.1186/s13148-020-00864-y
Frequently asked questions
General FAQs
- What are the SCAN Dashboards?
About the SCAN Public Dashboard
The SCAN Public Dashboard tracks the number of participants with SCAN-compliant MRI and PET images and the total number of images submitted by each Alzheimer’s Disease Research Center (ADRC) to SCAN that have passed QC. This resource is meant to provide ADRCs and researchers with up-to-date information on the number of standard images that are available for analysis. View a demo of the SCAN Public Dashboard.
Information included in the Dashboard:
Links to each center's ADRC-specific SCAN QC Dashboard
Total number of MRI and PET participants for whom the site has submitted data to SCAN
Total number of standard MRI exams and series submitted to SCAN that have passed QC and are available for analysis
Total number of PET scans (amyloid or tau) submitted to SCAN that have passed QC and are available for analysis
Information NOT included in the Public Dashboard:
SCAN analysis data for ADRCs
SCAN analysis data for Researchers: ADRD researchers around the world may access this data via NACC's Quick Access File Data Request System.
ADRC-specific PET and MRI data: The MRI and PET summary and QC status information is available through the ADRC-specific SCAN QC dashboards
Data for participants without existing NACCIDs: For the time being SCAN/LONI will only be accepting data submissions for participants who have existing NACCIDs
About the ADRC-specific SCAN QC Dashboards
Each ADRC can access their ADRC-specific SCAN QC dashboards via the SCAN Public Dashboard located above. The ADRC-specific SCAN QC dashboards will:
Enable them to track the QC status of their MRI and PET submissions, allowing them to rapidly address errors.
Serve as an auditable record of all SCAN MRI data submitted to SCAN by a given ADRC, downloadable as a csv file.
IMPORTANT NOTE: The ADRC-specific SCAN QC Dashboards are only available to ADRC members.
Information that will be included in the ADRC-specific SCAN dashboards includes:
All PET summary and QC data submitted by each ADRC including:
NACCIDs
PTIDs
PET study date and time
Image UID & Scanner model
PET radiotracer used
Whether the scan passed QC or not
QC comments (why the scan failed QC, if failed)
All MRI summary and QC data submitted by each ADRC including:
NACCIDs
PTIDs
MRI study date and time
Image UID & Scanner model
Series date and time
Series number
Series type
Study comments: Visual artifacts, etc.
Release for Analysis: Whether the series was released for analysis or not based on QC and protocol checks
The ADRC-specific SCAN QC Dashboards, available via the ADRC Portals, now include the numerical analysis results – such as Standardized Uptake Value Ratio ‘SUVRs’, brain volumes, white matter hyperintensities (WMHs), cortical thickness, surface area, and cerebral infarction– for all participants whose MR and PET images have passed QC and been analyzed.
- What is the current process for submitting MRI/PET images?
As of March 17, 2023, sites will only be able to submit SCAN MRI and PET images to LONI for participants that already have a NACCID. This change will prevent errors (i.e., duplicate NACC IDs for participants, etc.) from being introduced into the database and the need for future SCAN data audits. See Changes to the LONI Uploader section for more information.
Please note that this is a temporary change. Sites are encouraged to continue collecting SCAN data on all of their participants but will not be able to submit data for participants without existing NACC IDs until a new process for assigning NACCIDs to SCAN participants is put in place. NACC hopes to provide further updates to ADRCs on this new process soon.What about non-SCAN compliant MRI/PET images that are submitted to NACC?
As of March 17, 2023, sites will only be able to submit non-standard (non-SCAN compliant) MRI and PET images to NACC for participants that already have pre-existing NACCIDs.
Please do not submit non-standard MRI and PET images to SCAN as the SCAN initiative only collects SCAN-compliant MRI and PET images.Existing MRI and PET images collected before January 1, 2021, and prospective images that don't meet SCAN protocol are still accepted by NACC.
Learn more about submitting this data to NACC here. - How are MR/PET image submissions being tracked?
SCAN-compliant MRI/PET images submitted to LONI as part of the SCAN initiative:
SCAN Public Dashboard:The Public SCAN Dashboard tracks the total number of SCAN MRI and PET data submitted by each ADRC that have passed QC and are available for analysis. The totals reflected in the SCAN Public Dashboard differ from the total number of images your ADRC has submitted as it takes time for the SCAN team to evaluate images for compliance and to perform de-facing, quality control, and analysis prior to releasing this data to researchers. For a more complete picture of your total submissions by center and where they are at in the QC pipeline, please visit your ADRC-specific SCAN QC Dashboard via the “private dash” linked below.
ADRC-Specific SCAN QC Dashboards As part of the ADRC Portals, each ADRC has access to their ADRC-specific SCAN QC dashboards that a) enable them to track the summary and QC status of their MRI and PET submissions, allowing them to rapidly address errors, and b) serve as an auditable record of all SCAN MRI data submitted to LONI/NACC by a given ADRC, downloadable as a csv file.
Sites also can access the analysis results for their participants within the ADRC Portals in addition to SCAN documentation and resources.
The NACC monthly reports for each ADRC do not include the MRI/PET images submitted to LONI for the SCAN project in the image counts. We will not be adding SCAN images to these static monthly reports because we are in the process of modernizing the NACC Data Platform and complete SCAN-compliant image counts will be available through the SCAN dashboards.
Non-SCAN compliant MRI/PET images submitted to NACC:
NACC is currently in discussion with NIA and the Imaging Steering Committee to determine how mixed protocol (previously `legacy`) MRI and PET images will be tracked and reported at NACC as we transition to our modern Data Platform and ADRC portals.
Your monthly report should currently reflect the correct number of mixed protocol (previously `legacy`) MRI and PET images that have been submitted to by your site to NACC.
If you are concerned that the image counts in the report for your center do not accurately reflect the mixed protocol images your center has submitted to NACC, please reach out to nacchelp@uw.edu.
- How will MR/PET image submissions be tracked in the future?As NACC completes its transition onto our new modern data platform, we will launch the following resources:
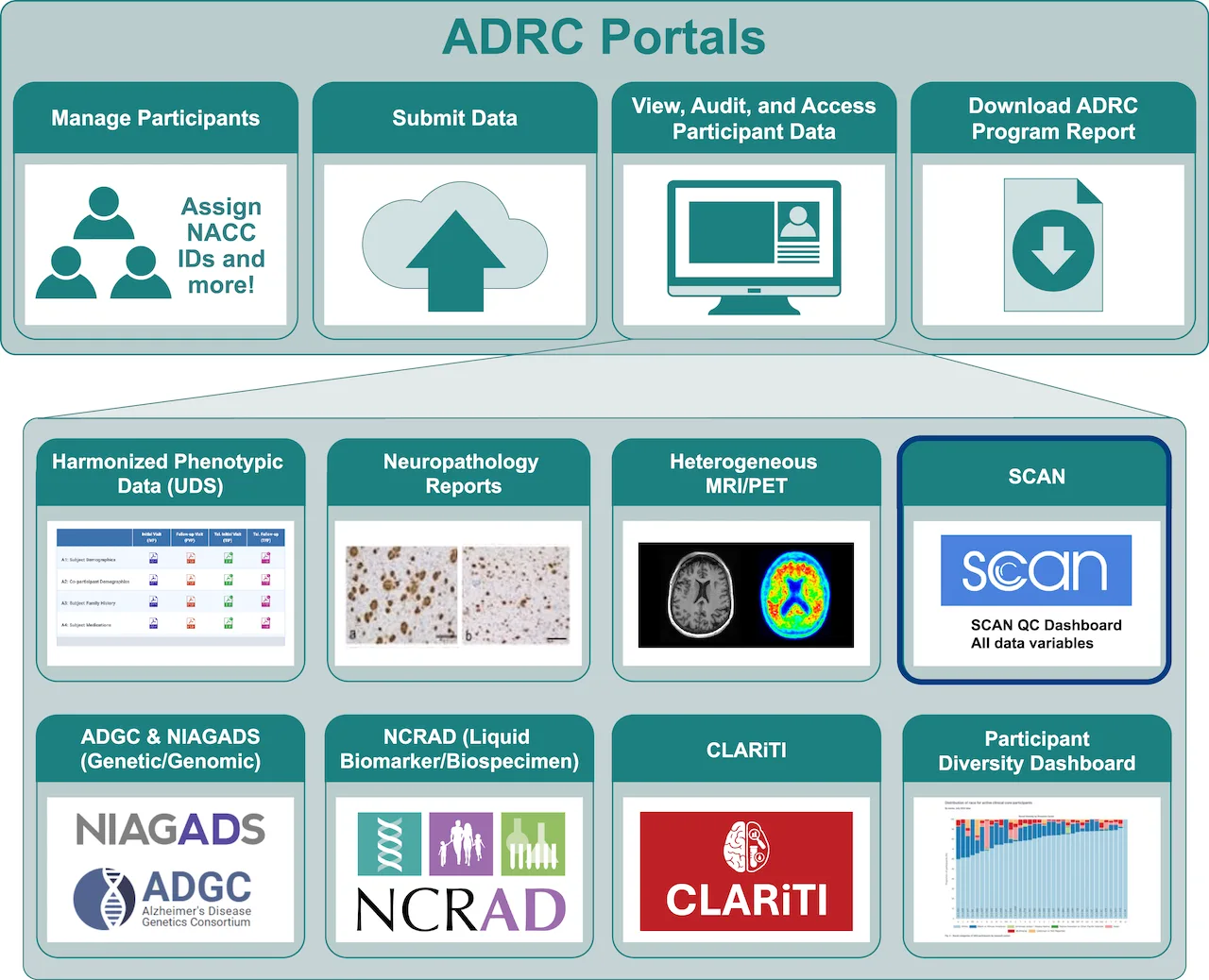
ADRC Portals
The ADRC Portals will provide ADRCs with self-service access to view, download, and audit their real-time data. These portals will replace inefficient static monthly reports with modern real-time ADRC-specific data portals (ADRC Portals), fulfilling an emergent request from ADRCs to have better access to their data submissions.
The ADRC Portals will provide ADRCs with access to six core data/metadata types:
UDS
Neuropathology
Non-standard MRI/PET
Standard MRI/PET
Biospecimen (NCRAD) and Genetic/genomic (NIAGADS)
Participant Diversity Metrics
NACC is currently working to build the ADRC Portals within NACC's new Data Platform (powered by Flywheel). Flywheel will support the need to make this data accessible only to designated ADRC representatives, requiring management by ADRC personnel and the technology platform to allow access control for appropriate viewers. NACC is using user centered design to develop dashboards to adapt the Flywheel interface to the ADRC programs' unique needs.
NACC's new Data Platform
The NACC Data Platform (NACC DP) that will integrate a host of existing and new standardized and longitudinal data streams from across the ADRC Program as well as metadata and analysis from partner organizations and initiatives, with all data streams linked by NACC Identifiers (NACC IDs). The NACC DP will include a variety of data search and access interfaces including the Data Front Door, ADRC Data Portals, and the NIA Portal.
These systems have been designed with an eye towards needing a modern tools that will not only accommodate our existing long-standing data streams such as the Uniform Data Set (UDS), neuropathology reports, and non-standard MRI/PET but that will also scale to accommodate and share innovative new data streams such as standardized imaging data, digital neuropathology, and digital biomarkers, and that can integrate complementary metadata and analysis data for standardized MRI/PET imaging (SCAN/LONI), genetic/genomic (NIAGADS) and biomarker (NCRAD) data from other ADRD data repositories, with all data streams linked by NACC unique identifiers (NACC IDs).
The transition to this new data platform will begin by giving access to ADRC portals in which each center will be able to access their data and view dashboards allowing them to track their contributions to the ADRC program. NACC will increasingly add functionality to these portals, ultimately replacing old processes like emailed monthly reports.
- How are identifiers used for SCAN data?
When submitting MRI/PET images to SCAN via the LONI uploader, you are currently asked to enter the Subject ID or PTID. This number is different from the NACCID as described below:
Subject ID or PTID: this is the ADRC-managed participant ID, or "local ID". Formats vary by ADRC.
NACCID: this is the NACC-managed participant ID. It is a string with the prefix 'NACC' followed by 6 digits.
ADCID: this is the NACC-managed center ID. This is a two-digit number.
LONIUID: this is the unique ID assigned to your images upon upload at LONI.
Each PTID and ADCID pair maps to a unique NACCID.
- How will ADRCs contributing data to SCAN be acknowledged?
NACC will build off the current NACC Data Use Agreement (DUA) policy to provide researchers with a comprehensive list of ADRCs and other related grants who have contributed their data to SCAN. All researchers and person(s) requesting data from NACC (including SCAN data in the future) must accept the NACC DUA policy.
The current NACC DUA policy requires authors to include the following statement acknowledging the NACC grant and Center support in presentations and the acknowledgments section of manuscripts:
The NACC database is funded by NIA/NIH Grant U24 AG072122. NACC data are contributed by the NIA-funded ADRCs: P30 AG...., etc.
If your site is involved in the SCAN initiative and you have additional grants or updates to include in the acknowledgements, please reach out to naccmail@uw.edu.
FAQs for ADRCs participating in SCAN
- Is my MRI/PET data SCAN-compliant (based on Human Subjects regulations/IRB)?You must acquire images under an IRB-approved consent that allows sharing of de-identified data to contribute this data to SCAN. It is the responsibility of each ADRC to ensure their consents allow them to share de-identified data, or to update study consents (and obtain IRB approval) to allow this type of sharing on prospective data.
- Is my MRI/PET data SCAN-compliant (based on NACC participation and acquisition parameters)?SCAN images must come from NACC participants. Details on allowed acquisition parameters are available in SCAN MRI manual and the SCAN PET manual.
- Our PET data are acquired with slightly different timing and/or reconstruction parameters than listed in the SCAN PET technical manual – can we still contribute this data to SCAN?No. Please only submit PET images that follow allowed acquisition parameters; submit all other images to NACC directly (learn more here).
- How do I upload my MRI/PET data to SCAN?Upload SCAN image data and corresponding upload forms to LONI following instructions detailed in the SCAN MRI and SCAN PET manuals.
- Why do I need to provide funding source when uploading images?As of August 9, 2025, ADRC’s will be required to provide Funding Source for each scan as part of the MRI and PET image upload process via the LONI IDA uploader. This additional information is critical to helping track ADRC contributions to their P30 imaging requirements as per the updated ADRC P30 RFA requirements which detail that ADRCs must collect and submit “Complete Magnetic Resonance Imaging, Amyloid positron emission topography (PET) Imaging, and Tau PET Imaging in at least 24 participants of the Clinical Core each year. All three imaging modalities must be completed for each participant. The Administrator is to ensure the minimum is met and distribute the funds appropriately to the relevant component(s) to accomplish the goals, including transmission of the data to SCAN.”
- How can I track my ADRC’s progress on the P30 required submissions?Sites are responsible for tracking and reporting on their P30 funding requirements. To assist in this process, the MRI and PET scan funding source metrics will be made available to ADRCs, coming soon: The SCAN QC Dashboards in the ADRC Portals will allow you to track the number of SCAN image submissions that meet the P30 RFA requirements. The CLARiTI Imaging Dashboards in the ADRC Portals will allow you to track the number of CLARiTI image submissions that meet the P30 RFA requirements. Please note that these metrics are not currently available but will be made available soon.
- Which participant ID do I use when uploading images?Users must submit the participant ID (also called PTID, Subject ID, or Local ID) and the participant’s NACCID. If there is a mismatch, then the user will receive an error message indicating that the PTID and NACCID combination are not valid, and the upload will not continue.
- How do I get a SCAN user account on LONI?
- Why are all SCAN images required to be acquired January 2021 or later?SCAN is a prospective project that officially launched January 2021. SCAN will not ingest data from before 2021 as retrospective or legacy data were not included in the project aims. There are other, separately funded projects ongoing to harmonize legacy MRI and PET data available in NACC, and SCAN will work with these investigators to help make their efforts compatible with SCAN data.
Contact Us
SCAN PIs
Bill Jagustjagust@berkeley.edu
Cliff Jackjack.clifford@mayo.edu
SCAN PET Team
Tessa Harrisontessaharrison@berkeley.edu
SCAN MRI TeamSCANMRI@mayo.edu
Office Hours
The first Tuesday of every month we will hold open office hours via Zoom. SCAN investigators are standing by and prepared to answer your questions about imaging protocols, uploading, private imaging dashboards or any SCAN (including CLARiTI images, and P30-funded images) related issues you have.







